Difference between revisions of "HMM and alignment"
Devicerandom (Talk | contribs) |
Devicerandom (Talk | contribs) |
||
| Line 1: | Line 1: | ||
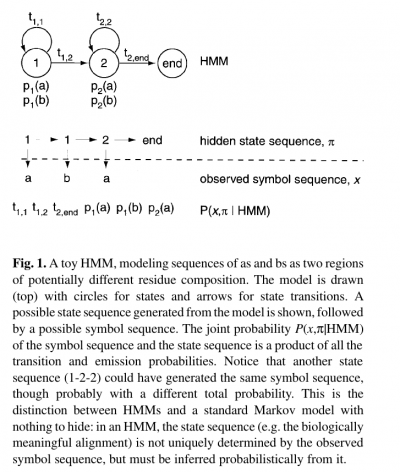
| + | * The name ‘hidden Markov model’ comes from the fact that the state sequence is a first-order Markov chain, but only the symbol sequence is directly observed. | ||
* Profile HMMs are similar to simple sequence profiles, but in addition to the amino acid frequencies in the columns of a multiple sequence alignment they contain the position-specific probabilities for inserts and deletions along the alignment | * Profile HMMs are similar to simple sequence profiles, but in addition to the amino acid frequencies in the columns of a multiple sequence alignment they contain the position-specific probabilities for inserts and deletions along the alignment | ||
* The logarithms of these probabilities are in fact equivalent to position-specific gap penalties (Durbin et al., 1998). | * The logarithms of these probabilities are in fact equivalent to position-specific gap penalties (Durbin et al., 1998). | ||
[[File:HMM 1998 review.png|400px]] | [[File:HMM 1998 review.png|400px]] | ||
| + | |||
| + | |||
| + | * The states of the HMM are often associated with meaningful biological labels, such as ‘structural position 42’. In our toy HMM, for instance, states 1 and 2 correspond to a biological notion of two sequence regions with differing residue composition. | ||
| + | * Inferring the alignment of the observed protein or DNA sequence to the hidden state sequence is like labeling the sequence with relevant biological information. | ||
| + | |||
* '''The alignment algorithm maximizes a weighted form of coemission probability, the probability that the two HMMs will emit the same sequence of residues. ''' | * '''The alignment algorithm maximizes a weighted form of coemission probability, the probability that the two HMMs will emit the same sequence of residues. ''' | ||
Revision as of 18:22, 11 February 2013
- The name ‘hidden Markov model’ comes from the fact that the state sequence is a first-order Markov chain, but only the symbol sequence is directly observed.
- Profile HMMs are similar to simple sequence profiles, but in addition to the amino acid frequencies in the columns of a multiple sequence alignment they contain the position-specific probabilities for inserts and deletions along the alignment
- The logarithms of these probabilities are in fact equivalent to position-specific gap penalties (Durbin et al., 1998).
- The states of the HMM are often associated with meaningful biological labels, such as ‘structural position 42’. In our toy HMM, for instance, states 1 and 2 correspond to a biological notion of two sequence regions with differing residue composition.
- Inferring the alignment of the observed protein or DNA sequence to the hidden state sequence is like labeling the sequence with relevant biological information.
- The alignment algorithm maximizes a weighted form of coemission probability, the probability that the two HMMs will emit the same sequence of residues.
- Amino acids are weighted according to their abundance, rare coemitted amino acids contributing more to the alignment score.
- Secondary structure can be included in the HMM-HMM comparison.
- We score pairs of aligned secondary structure states in a way analogous to the classical amino acids substitution matrices.
- We use ten different substitution matrices that we derived from a statistical analysis of the structure database, one for each confidence value given by PSIPRED.