Difference between revisions of "HMM and alignment"
Devicerandom (Talk | contribs) (→How it works inside) |
Devicerandom (Talk | contribs) |
||
| Line 47: | Line 47: | ||
| − | [[Category:Bioinfo notes]] | + | [[Category:Bioinfo notes|Hidden Markov models]] |
Revision as of 19:03, 11 February 2013
This is a collage of stuff from papers and HHpred website
Building a profile HMM
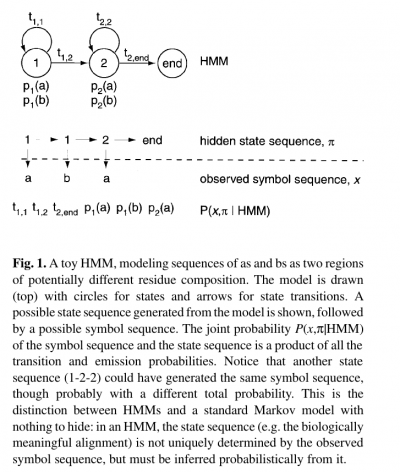
- The name ‘hidden Markov model’ comes from the fact that the state sequence is a first-order Markov chain, but only the symbol sequence is directly observed.
- Alternatively, an HMM can be built from prealigned (pre-labeled) sequences (i.e. where the state paths are assumed to be known).
- In the latter case, the parameter estimation problem is simply a matter of converting observed counts of symbol emissions and state transitions into probabilities.
- In building a profile HMM, an existing multiple alignment is given as input.
- it pays to build HMMs on pre-aligned data whenever possible.
- Especially for complicated HMMs, the parameter space may be complex, with many spurious local optima that can trap a training algorithm.
How it works inside
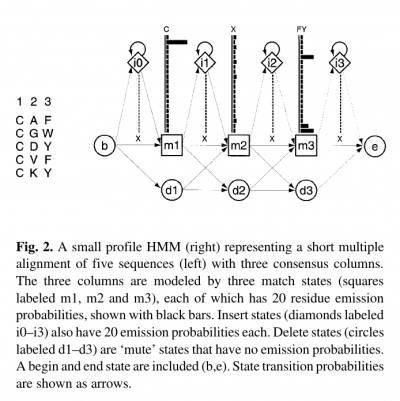
- Profile HMMs are similar to simple sequence profiles, but in addition to the amino acid frequencies in the columns of a multiple sequence alignment they contain the position-specific probabilities for inserts and deletions along the alignment
- For each consensus column of the multiple alignment, a ‘match’ state models the distribution of residues allowed in the column.
- An ‘insert’ state and ‘delete’ state at each column allow for insertion of one or more residues between that column and the next, or for deleting the consensus residue.
- Profile HMMs are strongly linear, left right models, unlike the general HMM case.
- The probability parameters in a profile HMM are usually converted to additive log-odds scores before aligning and scoring a query sequence (Barrett et al., 1997).
- The scores for aligning a residue to a profile match state are therefore comparable to the derivation of BLAST or FASTA scores: if the probability of the match state emitting residue x is px , and the expected background frequency of residue x in the sequence database is fx , the score for residue x at this match state is log(px/fx).
- The logarithms of these probabilities are in fact equivalent to position-specific gap penalties (Durbin et al., 1998).
- In contrast to parameter estimation, a suitable HMM architecture (the number of states, and how they are connected by state transitions) must usually be designed by hand.
- A maximum likelihood architecture construction algorithm exists for the special case of building profile HMMs from multiple alignments (Durbin et al., 1998).
- Insertions tend to be seen most often in surface loops of protein structures, and so have a bias towards hydrophilic residues.
- Profile HMMs can capture this information in the insert state emission distributions.
Meaning of HMM states
- The states of the HMM are often associated with meaningful biological labels, such as ‘structural position 42’. In our toy HMM, for instance, states 1 and 2 correspond to a biological notion of two sequence regions with differing residue composition.
- Inferring the alignment of the observed protein or DNA sequence to the hidden state sequence is like labeling the sequence with relevant biological information.
Alignment of HMMs
- The alignment algorithm maximizes a weighted form of coemission probability, the probability that the two HMMs will emit the same sequence of residues.
- Amino acids are weighted according to their abundance, rare coemitted amino acids contributing more to the alignment score.
- Secondary structure can be included in the HMM-HMM comparison.
- We score pairs of aligned secondary structure states in a way analogous to the classical amino acids substitution matrices.
- We use ten different substitution matrices that we derived from a statistical analysis of the structure database, one for each confidence value given by PSIPRED.